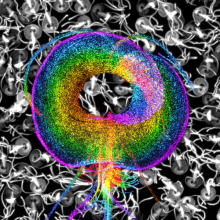
Recently developed tissue-hydrogel methods for specimen expansion now enable researchers to perform super-resolution microscopy with ∼65 nm lateral resolution using ordinary microscopes, standard fluorescent probes, and inexpensive reagents. Here we use the combination of specimen expansion and the optical super-resolution microscopy technique structured illumination microscopy (SIM) to extend the spatial resolution to ∼30 nm. We apply this hybrid method, which we call ExSIM, to study the cytoskeleton of the important human pathogen Giardia lamblia including the adhesive disc and flagellar axonemes. We determined the localization of two recently identified disc-associated proteins, including DAP86676, which localizes to disc microribbons, and the functionally unknown DAP16263, which primarily localizes to dorsal microtubules of the disc overlap zone and the paraflagellar rod of ventral axonemes. Based on its strong performance in revealing known and unknown details of the ultrastructure of Giardia, we find that ExSIM is a simple, rapid, and powerful super-resolution method for the study of fixed specimens, and it should be broadly applicable to other biological systems of interest.